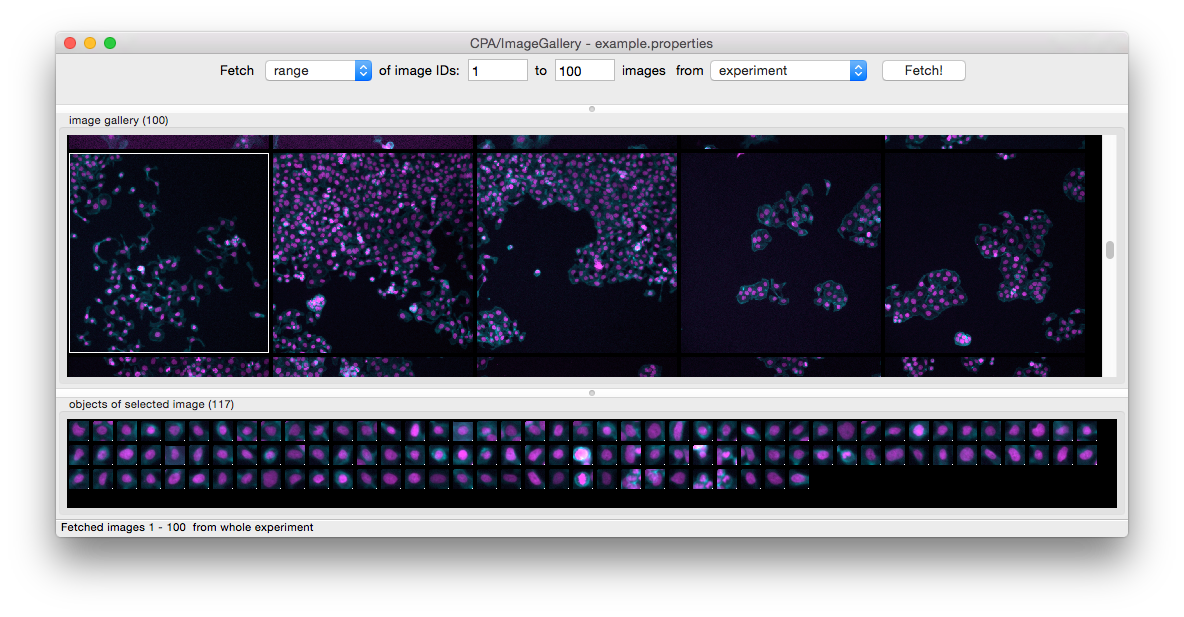
c) Execution time to run the training sequence for a FastGentleBoosting classifier. b) Comparison of execution time between CPA 2.2 and CPA 3.0 when loading a training set of 600 objects in 13 classes. 
The lasso tool has been activated to select objects of interest, which can then be used to train a classifier. Marker colors represent classification results from a previously generated classifier. a) Screenshot of the new Dimensionality Reduction tool, using principal component analysis with the example dataset. Improvements in CellProfiler Analyst 3.0. This provides a more manageable series of features which the user can then explore. Dimensionality reduction allows the numerous measurements generated by CellProfiler to be condensed into a smaller subset of features which represent the overall variance in the dataset. When the dataset contains thousands of measurements users can struggle to identify those which reveal these outliers. This is particularly important when creating classifiers that can identify rare images in a set rare outliers are needed for training but may be difficult to find through random sampling. We added a dimensionality reduction tool to help users visualize the variance within high-dimensional datasets ( Fig. 1A). This maintains and expands upon the interoperability between the two programs. At present neither of these loaders supports accessing only partial sections of an image, which may be an area for further development to handle large files produced by whole slide imaging.ĬellProfiler Analyst 3.0 can export machine learning models that are compatible with CellProfiler 4.2+, allowing the resulting classifiers to be directly embedded into CellProfiler pipelines. While bioformats provides broader file format compatibility, imageio allows for common image formats to be loaded more efficiently, which greatly improves the time to generate object thumbnails. In addition, we added a faster imageio-based image loader to supplement the existing bioformats-based implementation ( Silvester et al., 2020). We also revised the program’s builds to package all Java dependencies within the main installer, which dramatically simplifies the installation process. We ported CellProfiler Analyst to the Python 3 programming language to ensure compatibility with future operating systems after the official Python 2 end-of-life in 2020. Herein we present CellProfiler Analyst 3.0, which includes major performance improvements and new features that improve the utility of the software. These tools provide an intuitive interface for scientists to explore their data in forms such as histograms and scatter plots, though without the advanced statistical tools or extreme customizability of a pure programming language. While other tools such as Ilastik ( Berg et al., 2019) and Advanced Cell Classifier ( Piccinini et al., 2017) can provide a GUI for training classifiers with image-based data, these do not include data exploration and visualization tools like those in CellProfiler Analyst. The software includes several tools for users to visualize and filter their datasets, alongside tools for training machine learning classifier models within a convenient graphical user interface that is geared toward working with image data ( Dao et al., 2016). 
CellProfiler Analyst is a data exploration package for helping users to explore and extract information from large datasets, including (but not limited to) those produced by CellProfiler pipelines ( Jones et al., 2008). Spreadsheet programs are familiar but lack features and integration with source images that biologists often need. Free software packages such as ImageJ ( Schneider et al., 2012) and CellProfiler ( McQuin et al., 2018) allow users to extract hundreds or thousands of numerical measurements from their image data, but it is ultimately on the user to determine which of these features are relevant to the biological problem being investigated. This necessitates automated computational analysis to efficiently derive biological insights from the raw data. With the increasing adoption of high-throughput microscopy, scientists have been able to generate large datasets containing thousands of individual images.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
